# A tibble: 2 × 3
arms_reduction num_responses pct
<fct> <int> <dbl>
1 against 452 0.440
2 favor 576 0.560Inference for a single proportion
IMS, Ch. 16
Smith College
Apr 1, 2026
Inference for a single proportion
Nuclear Arms Reduction Survey
A simple random sample of 1,028 US adults in March 2013 found that 56% support nuclear arms reduction.
- Does a majority support nuclear arms reduction?
Sample proportion (\(\hat{p}\))
Response: arms_reduction (factor)
# A tibble: 1 × 1
stat
<dbl>
1 0.560- Note:
p_hatis a \(1 \times 1\)data.frame p_hat$statis the actual (scalar) value
NHST Setup
- \(H_0: p = 0.5\)
- \(H_A: p \neq 0.5\)
- \(\alpha = 0.05\)
Big Question
How do we construct the null distribution?
Three methods for inference
- Use probability theory
- today
- Use normal approximation
- also today
- mathematical model (Ch. 16.2)
- Use simulations
- parametric bootstrap (Ch. 16.1)
What do we need to know about the null dist?
- Center, shape, and spread!
- Once we have the null distribution…
- What is the standard error?
- Where does the test statistic lie in the null distribution?
- What is the p-value?
- What is our decision?
Method 3: Simulation
Parametric bootstrap (review)
- Draw samples from binomial distribution
Null distribution (simluated)
Null distribution (simulated, in black)
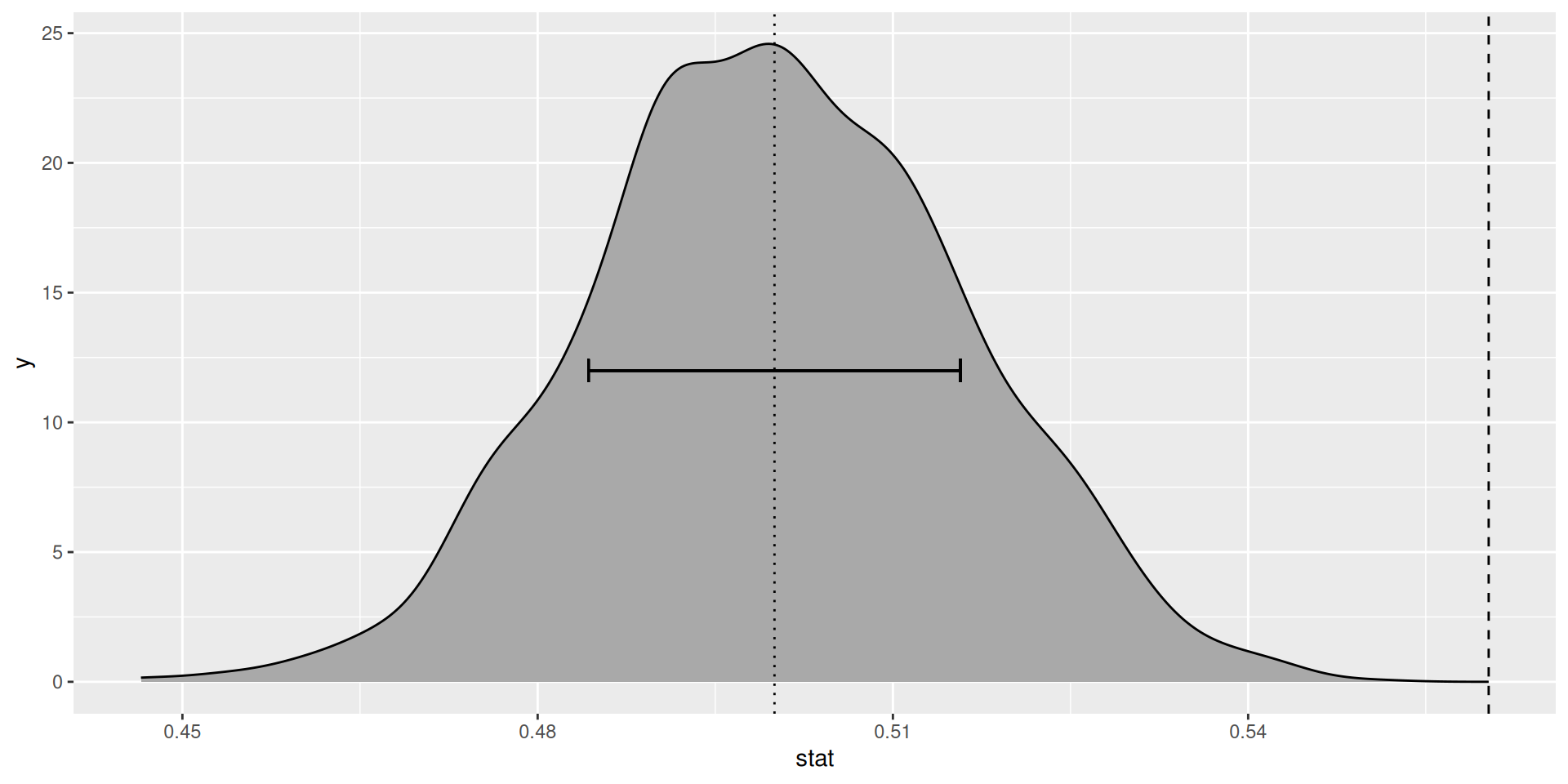
Standard error and p-value
- Standard error (\(SE_{\hat{p}}\)):
Your turn
- Does a majority support nuclear arms reduction?
Why simulate the null?
Pros:
- intuitive
- no math
- few assumptions
Cons:
- requires computer
- requires coding ability
- solution is approximate
- non-deterministic
Method 1: Probability theory
Probability theory
- Compute exact null distribution
Let \(X \sim Bernoulli(p)\). Then,
\[ \mathbb{E}[X] = p, Var(X) = p(1-p) \]
Let \(Y = X_1 + \cdots + X_n\). Then \(Y \sim Binom(n, p)\) and,
\[ \mathbb{E}[Y] = np, Var(Y) = np(1-p) \]
Probability theory (cont’d)
\(Z = Y/n\) is a r.v. giving the mean of \(n\) draws from \(X\)!
Then,
\[ \mathbb{E}[Z] = p \]
And,
\[ Var(Z) = \frac{1}{n^2} \cdot np(1-p) = \frac{p(1-p)}{n} \]
And, \(sd(Z) = \sqrt{\frac{p(1-p)}{n}}\) is the standard error!
Properties of null distribution
- Same shape as binomial distribution curve
- In our case, \(\hat{p} \sim Z\) and \(\mathbb{E}[X_i] = p_0\)
- Thus, \(SE_{\hat{p}} = \sqrt{\frac{p_0(1-p_0)}{n}}\)
se <- sqrt(p_0 * (1 - p_0) / n)
dbinom_p <- function (x, size, prob, log = FALSE) {
size * dbinom(round(x * size), size, prob, log)
}
null_dist_2 <- null_dist +
geom_function(
fun = dbinom_p, color = "red",
args = list(size = n, prob = p_0)
) +
geom_errorbarh(
aes(xmax = p_0 + se, xmin = p_0 - se, y = 10),
color = "red"
)Null distribution (exact, in red)
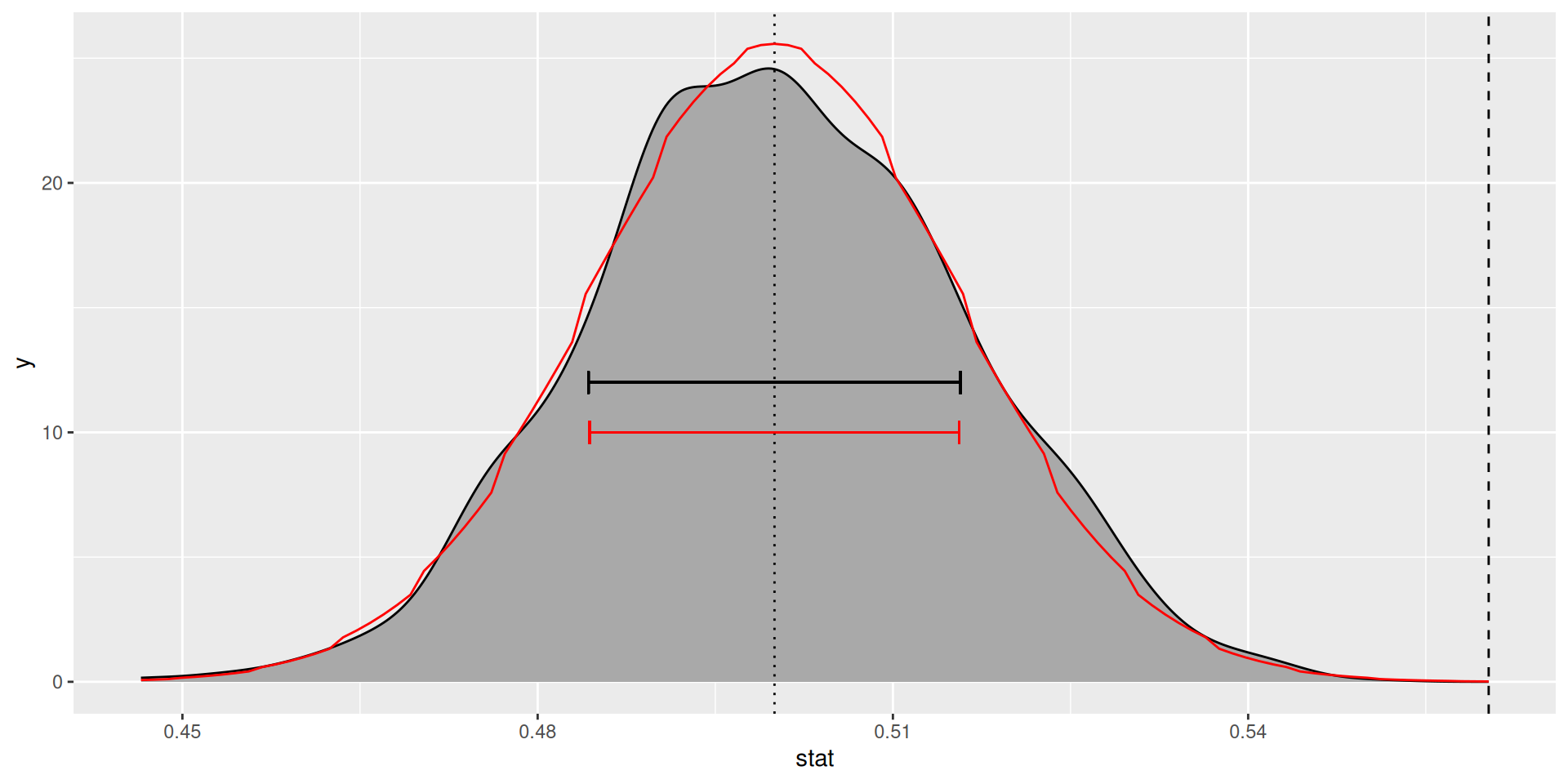
Standard error and p-value
- Standard error (\(SE_{\hat{p}}\)):
- p-value:
Your turn
Does a majority support nuclear arms reduction?
Compare the SE and p-value from the previous method. Are they meaningfully different?
Why compute the null?
Pros:
- correct answer
- deterministic
Cons:
- Had to do a bunch of math. Math wasn’t too bad in this case, but it gets much harder in other cases.
- only possible for simple situations
- For large \(n\), computation of binomial distribution is not that easy
Method 2: Normal approximation
Normal approximation
By CLT, for \(n\) large enough, null distribution is approximately normal
If \(np_0(1-p_0) > 10\), then approximation is reasonably good
Use \(SE_{\hat{p}} = \sqrt{\frac{p_0(1-p_0)}{n}}\) (from before)
- 💡💡 Now you know why!! 💡💡
Null distribution (normal approx.)
Null distribution (approx., in blue)
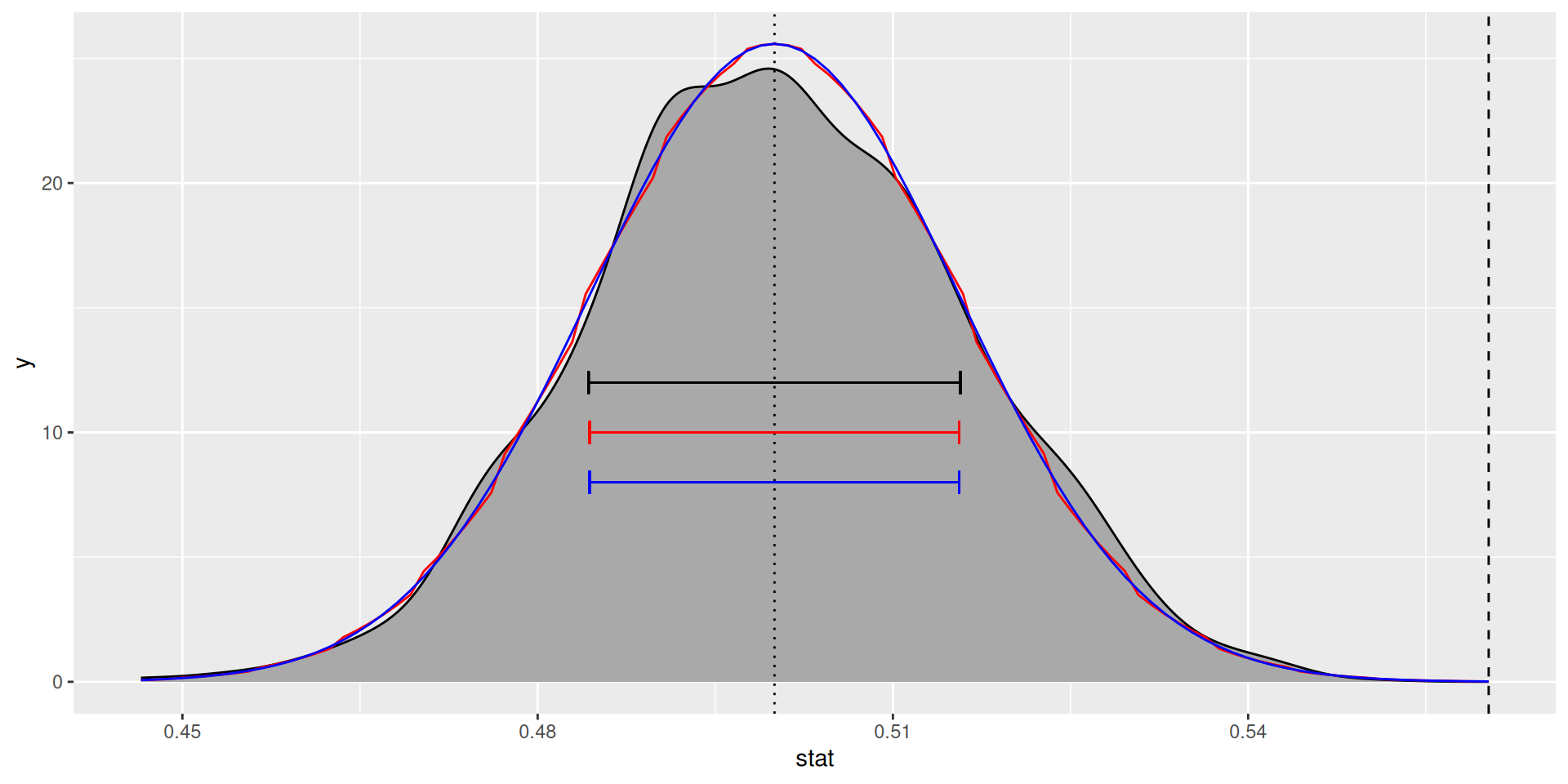
Standard error and p-value
- Standard error (\(SE_{\hat{p}}\)):
- p-value:
Z-scores
- Compute z-score:
Alternatively, using infer
Your turn
Does a majority support nuclear arms reduction?
Compare the SE and p-value from the previous method. Are they meaningfully different?
Why approximate the null?
Pros:
- provable approximation quality
- no computer required
- (kind of)
- deterministic
- widely-used
Cons:
Solution is approximate, and sometimes approximation is not great
SE formula comes out of nowhere
harder to connect to big ideas?
- Most common, well-understood method
